High Dimension To Low Dimension
April 13, 2019
高维数据映射为低维数据
高 —> 低
\[X_k = X \cdot W_k^T\]其中 𝑋𝑘 为映射后的低维数据,X 为原始的高维数据, Wk 为前 k 个主成分
低 —> 高
\[X_m = X_k \cdot W_k\]其中 𝑋m 为映射后的高维数据,Xk 为原始的高维数据映射后的低维数据, Wk 为前 k 个主成分,Xm 和原始 X 相比有损失
import numpy as np
import matplotlib.pyplot as plt
X = np.empty((100,2))
X[:,0] = np.random.uniform(0., 100., size=100)
X[:,1] = 0.75 * X[:,0] + 3. + np.random.normal(0, 10., size=100)
import numpy as np
class PCA:
def __init__(self, n_components):
assert n_components >= 1, "n_components must be valid"
self.n_components = n_components
self.components_ = None
def __repr__(self):
return "PCA(n_components=%d)" % self.n_components
def fit(self, X, eta=0.01, n_iters=1e4):
assert self.n_components <= X.shape[1],\
"n_components must not greater than feature number of X"
def demean(X):
return X - np.mean(X, axis=0)
def f(w, X):
return np.sum(X.dot(w)**2) / len(X)
def df(w, X):
return X.T.dot(X.dot(w)) * 2. / len(X)
def direction(w):
return w / np.linalg.norm(w)
def first_components(X, initial_w, eta, n_iters=1e4, epsilon=1e-8):
w = direction(initial_w)
cur_iter = 0
while cur_iter < n_iters:
gradient = df(w, X)
last_w = w
w = w + eta * gradient
w = direction(w)
if(abs(f(w,X) - f(last_w,X)) < epsilon):
break
cur_iter += 1
return w
X_pca = demean(X)
self.components_ = np.empty(shape=(self.n_components, X.shape[1]))
for i in range(self.n_components):
initial_w = np.random.random(X_pca.shape[1])
w = first_components(X_pca, initial_w, eta, n_iters)
self.components_[i] = w
X_pca = X_pca - X_pca.dot(w).reshape(-1,1) * w
return self
def transform(self, X):
assert self.components_ is not None, "must fit before transform"
assert X.shape[1] == self.components_.shape[1]
return X.dot(self.components_.T)
def inverse_transform(self, X):
assert self.components_ is not None, "must fit before transform"
assert X.shape[1] == self.components_.shape[0]
return X.dot(self.components_)
pca = PCA(n_components=2)
pca.fit(X)
PCA(n_components=2)
pca.components_
array([[ 0.77871582, 0.62737682], [-0.62737329, 0.77871866]])
pca = PCA(n_components=1)
pca.fit(X)
PCA(n_components=1)
X_reduction = pca.transform(X)
X_reduction.shape
(100, 1)
X_restore = pca.inverse_transform(X_reduction)
X_restore.shape
(100, 2)
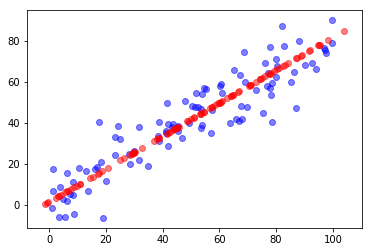
plt.scatter(X[:,0], X[:,1], color='b', alpha=0.5)
plt.scatter(X_restore[:,0], X_restore[:,1], color='r', alpha=0.5)
plt.show()
scikit-learn 中的 PCA
from sklearn.decomposition import PCA as SKPCA
skpca = SKPCA(n_components=1)
skpca.fit(X)
PCA(copy=True, iterated_power=’auto’, n_components=1, random_state=None, svd_solver=’auto’, tol=0.0, whiten=False)
skpca.components_
array([[0.77871588, 0.62737674]])
X_SKReduction = skpca.transform(X)
X_SKReduction.shape
(100, 1)
X_SKRestore = skpca.inverse_transform(X_SKReduction)
X_SKRestore.shape
(100, 2)
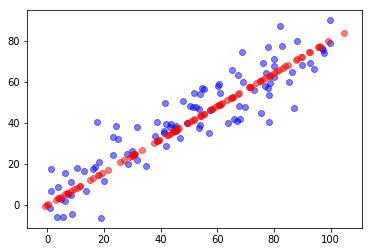
plt.scatter(X[:,0], X[:,1], color='b', alpha=0.5)
plt.scatter(X_SKRestore[:,0], X_SKRestore[:,1], color='r', alpha=0.5)
plt.show()